
INTRODUCTION
Most of the relationships we sought to prove are too many generations ago to give reliable DNA matches through conventional means. The potential link between for example the English and Barrow Lousadas and their 10th and 11th cousins among the USA Lousadas goes back 9 generations to different siblings of Amador de Lousada of Vinhais. Even 4th cousins do not show up well, for I have only one (25cM) segment in common with my 4th cousin Jeremy Lousada, and only 3 3cM matches with my 6th cousin John Griffiths. From the outset, we were well aware of the poor match-prediction powers of small (3cM) segment matches. Even at 7cM there is significant uncertainty as to their reliability, and only going to 15M gives this. Therefore we sought to rely on maximising the number of matches and then using statistical and related techniques to discern patterns lying within our datasets.
As Qmatch was recommended to us for small matches by GEDmatch, and retaining our desire to accumulate many matches, we set about using Qmatch (set at 3cM, P=3) to compile all 2255 segment matches - mostly off-target matches and false positives of course - from our 13 relative sample - see here the 1963 matches found before ELL was added to the sample of relatives. We found 46 RSBCs, 25 lefthand and 21 righthand, and most were new to us. We spent much time looking at RSBCs, how frequently they occur across chromosomes and in the match-rich areas on chromosomes. But they proved quite difficult to work with, and were relatively unproductive.
We then moved from RSBCs to ASBs; but they proved misleading and just as we prematurely claimed in 'Fun with Autosomal DNA', we again thought that we may have established a genetic link between the USA Lousadas, the English Lousadas, the Barrows, Scott's wife (and hence the Fischls) and Randy's parents. The problem with ASBs was that instead of an ASB being associated with a unique crossover, in fact many unrelated crossovers can all report to the same ASB (in fact the same applies to RSBCs as well). This is because a crossover is only measured as lying between 2 particular SNPs - these SNPs being the first non-matching SNP beyond the crossover at one end of a segment and last SNP within the segment at the other end of the segment. Typically there are 5000 base-pair positions between each SNP so the potential for unrelated and (from the family viewpoint) spurious crossovers at the same ASB is large. That is, ASBs are not the amazingly precise way of penetrating the fog of unreliable small matches that was hoped.
From Chart 1 we can see the futility of the 3cM P=3 results. For at this setting the randoms show more segment-matches than the relatives - 2261 to 2255 - that is, family does not show a signal! While the RSBC totals indicate that a family signal may exist, the ASB numbers seem odd (and perhaps can only be explained by a combination of spurious crossovers and the much greater possible ancestor pool where there is no genealogy).

CHART 1- COMPARISON DATA RELATIVES AND RANDOMS
AT LAST THE RELATIVES STAND OUT
From this impasse it was timely advice from GEDmatch in
February 2026 which assisted us. Thus, we came to use Qmatch to
look again at all 3cM segment-matches, not at P=3 but at P=7. These conditions give reasonable quality
segment-matches, almost as good as 7cM matches from some other providers. And
most importantly Chart 1 shows that at
3cM P=7, the relatives show 560 segment-matches which is 187 more
than the randoms' 373 segment-matches, the strongest such family signal we have seen in all our
comparisons of
relatives with randoms. Chart 2 shows the segment-match numbers for relative
pairs. For example, among the 14 B/Je segment-matches is one of 5cM on Cr10 which is
presumably their Ancestry.com 7cM match. JG misses matching (shown in grey) 2 of
the Barrow-Lousadas (A and J) whose specific ancestry evidently differs from
that of Ju.

CHART 2 - RELATIVES SEGMENT-MATCHES
COMPARISON OF RELATIVES AND RANDOMS
JG's 2 misses in Chart 2 contrast with 9 misses by the randoms in Chart 4. The presence of 5cM and 7cm segment-matches in Chart 4 is indicated by the same colours as in Chart 2. Further, we see that the randoms have 4 people in less than 4% of segment-matches compared with 1 relative (J, who despite proven close Lousada genetic and genealogical links, has anomalously low segment-match numbers), and 4 people in more than 10% of segment-matches (compared with 1 relative). That is, as expected, family connections despite some stochastic phenomena also being present, are tighter. In addition, the relatives show 9 triangulations compared with 2 shown by the randoms. Triangulations are discussed below, where it will be seen that 3 out of 9 are real for the relatives using the methodology we develop, but none of the 2 for the randoms.
But our first task is to assess how well our grouping of relatives (indicated by lines across the above table) compares with random. As there is only 1 English Lousada we can only consider 3 not 4 groupings here. We can see in Chart 3 that the intrabranch matching within each of the 3 groupings comfortably exceeds that of the random set. Supporting this we note that a triangulation is shown by the 1st and 4th cousins A, Ju and J at Cr16 (3219600 - 6259081) within the Barrow-Lousadas, and the B/Je/TP triangulation among the US Lousadas at Cr2 (217m - 220m).

CHART 3 - INTRABRANCH MATCHING
ANALYSIS OF RANDOMS
The random set could contain its own (but unexpected/unknown) internal family links which must be allowed for when we consider the real segment-match count and the false (off-target) segment-match count. Chart 4 shows the random set does indeed have structure with several obvious groupings. The central grouping seems to reflect several (perhaps 3) Ashkenasi families variously linked perhaps 4-8 generations ago. A second grouping is more distantly related perhaps with more diverse ancestry. The outer grouping is only remotely related. From this, we can make a rough estimate of the number of real segment-matches among the 373 in the random set - simply by adopting the central group's 154 segment-matches, leaving 219 false segment-matches. The estimate depends upon the false segment-matches arising in the central grouping balancing the real segment-matches arising in the outer groupings, something which we have judged reasonable but not proven.

CHART 4 - RANDOMS SEGMENT-MATCHES
IMPLICATION FOR RELATIVES
Now we must find how the relatives generate 219 false segment-matches. An easy way of doing this is shown here from which we see that deleting all white and green pairs except those showing 9 or more segment-matches is clearly indicated. We note that we now have a probability estimate covering the false segment-matches (namely 39% or 219/560), so that a pair with 9 segment-matches is tantamount to a proven match since 0.39**9 = 0.0002 or 0.02%. Accordingly, we can now present in Chart 5 our estimate of real relative matching. Here the yellow, green and blue colours show matches between relatives. The blue entries show 9-segment matches as just discussed. The yellows have been reduced by the known false TP 7cM match (see triangulation discussion below).
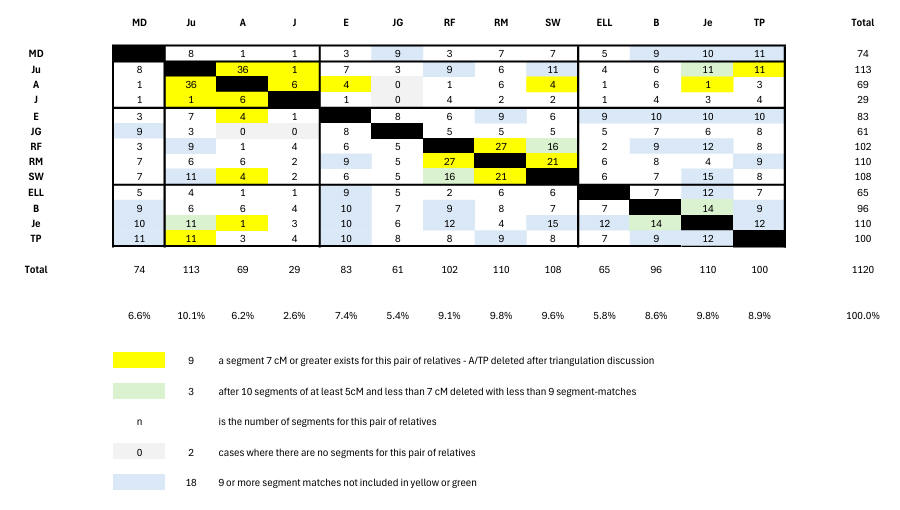
CHART 5 - RELATIVES REAL MATCHES
INTERBRANCH CONNECTION
There are 4 branches of relatives - English Lousada (MD), Barrow-Lousada (Ju, A, J), Barrow (E, JG, RM, RM, SW), and US Lousada (ELL, B, Je, TP). We extract from Chart 5 the number of matching relative-pairs for each of the 6 interbranch intersections, and express this as a percentage of total possible relative-pairs - see Chart 6. From this we can make some observations:
Most branches have matches with other branches - the exception being the sole English Lousada who does not show a match with the 3 Barrow Lousadas (despite a known Lousada connection!)
The English Lousada branch remarkably matches the Barrows and the US Lousadas despite no conceivable genetic connection other than the target Lousada link
The Barrow and US Lousada link is supported by the B/Je/E triangulation noted below.
The English Lousadas, the Barrows and the US Lousadas do best, achieving over 30% of possible matching pairs.
The Barrow Lousada branch does worst with 23% - despite its highest level of Lousada genes - due it seems to J and perhaps A being reluctant to show many segment-matches!

CHART 6 - EXTENT OF INTERBRANCH CONNECTIONS
TRIANGULATIONS
We find 9 triangulations among the relatives. We long knew of Ju/J/A on Cr 16, but there are 8 additional triangulations - namely Ju/RM/Je and RM/TP/Je and B/Je/TP and Ju/Je/B on Cr2 (217m - 220m), B/J/E on Cr5 (79m - 81m), J/TP/E on Cr10 (116m - 119m), A/E/RM on Cr17 (31m - 33m), and RM/B/Je on Cr22 (25640628 - 26225384). But at the Cr2 site only B/Je/TP is likely to be real being the only one having all 3 links from 9-segment pairs. The triangulations J/TP/E on Cr10 (116m - 119m), A/E/RM on Cr17 (31m - 33m), and RM/B/Je on Cr22 (25m - 26m) do not achieve this either, but the B/Je/E triangulation at Cr5 (79m - 81m) does. Thus apart from the intrabranch triangulations (Ju/J/A on Cr16 plus B/Je/TP on Cr2) we find one (interbranch) triangulation (B/Je/E on Cr5).
In summary, we have 3 real triangulations among relatives, one of which is interbranch with the other 2 being intrabranch.
The absence of a triangulation can also be informative - on Cr18 (6.6m - 8.1m) each of the 1st cousins Ju and A has a 7cM match with TP but don't match each other here which shows that at least one of the 2 matches is in fact false - probably A/TP. This of course shows that false matches can occur in the low-match yellow pairs, a possibility we allowed for in our discussion above.
CONCLUSION
We may feel confident that despite the small segment sizes and remote ancestral connection, there is considerable interbranch matching which supports our genealogy (in which the various branches originate in the ancestral family of Amador de Lousada).
FOOTNOTE
Finally, out of respect to those whose kit numbers went into generating the random sample, we comment further on Chart 4. This shows perhaps 3 family linkages around 4-8 generations ago (A/H/J/MW/DG) with a secondary group (N/S/M/P), together with an outlying group (C, MB, MMC, KB). The central 'family' group of 5 contains kits contributed from John Griffiths (A, H, J) and Julian Land (MW, DG). The secondary group of 4 consists of N contributed by John Griffiths and S, M, P contributed by Julian Land. The remotely-connected group of 4 consists of C contributed by John Griffiths and MB, MMC and KB contributed by Julian Land. The set of 13 randoms produced only 2 triangulations, on Cr1 at 158m - 160m (H, S, J) and on Cr8 at 11.3m - 12.6m (S, P, M). These triangulations mean little in this context, but despite this we note that only one of the 6 segment-matches comes from a pair having at least 9 segment-matches (S/P), meaning neither of the 2 triangulations in the random set would have been real in a family context.